[+]
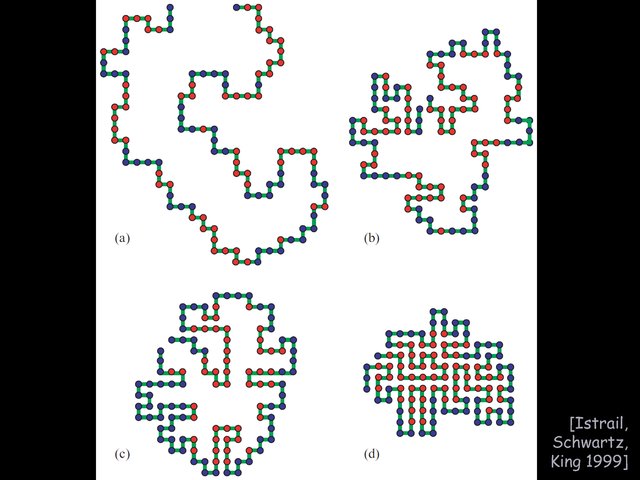
Protein folding: HP model of protein folding,
NP-hardness and approximation, unique optimal foldings, protein design.
Interlocked 3D chains: Lubiw's problem, unlocking 2-chains,
unlocking two 3-chains and 2-chains,
interlocking 3-chains, interlocking 3-chain with 4-chain,
interlocking 4-chain with triangle, interlocking 3-chain with quadrangle,
topological vs. geometric arguments, rigid and fixed-angle variations.
|
|
This lecture continues our discussion on protein folding,
this time focusing on simple theoretical models of the forces, rather
than the mechanics, behind protein folding. In particular, we'll
see the HP model, a lattice model capturing the hydrophobia of
certain amino, which try to hide from the surrounding water.
Finding the optimal folding a given protein is NP-complete, but
there are some decent constant-factor approximations, and it's also
not known whether protein design is similarly hard. We'll see one step
in the direction of design: guaranteeing a unique optimal folding.
Then we'll turn to a fun problem of interlocked linkages.
It's known that you need five bars to lock an open chain,
but can you interlock multiple chains each with less than five bars?
The answer is yes, with a nearly complete characterization of the
possibilities. One consequence we'll see is how to cut a chain of
n bars into around n/2 pieces that are not interlocked,
while n/4 pieces are necessary.
|